InitDF = pd.read_csv(file, sep="\t", header=0)įor chunks in pd.read_csv(file2, sep="\t", chunksize=50000, header=0): I know why its doing that but I'm not sure on how to fix this, right now my code for chunking is really simple. | 010 | CBA | CBA | 0 | 1 | 4 | 9 | 0 | NA | NA | NA | 0 | 0 | 1 |īasically it treats each chunk as if it were a new file and not from the same one. I already have code to change the names of the headers so I'm not worried about that, however when I use chunks I end up with something closer to - - - - - - - - - - - - - - When combining the two frames I need them to merge on "Mod", "Nuc" and "AA" so that I have something similar to this - - - - - - - - - - - My dataframes are similar to this (they can vary in size, but some columns remain the same "Mod", "AA" and "Nuc") - - - - - - - - My idea has been to load one dataframe into memory whole and then chunk the others and combine iteratively, this didn't work so well. 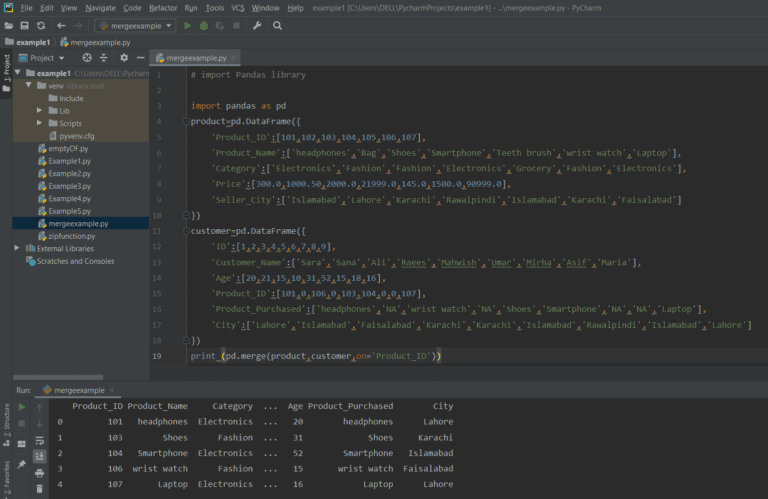
This is an issue for one reason, the script runs on an AWS server and running out of RAM means a server crash.Ĭurrently the file size limit is around 250mb each, and that limits us to 2 files, however as the company I work is in Biotech and we're using Genetic Sequencing files, the files we use can range in size from 17mb up to around 700mb depending on the experiment. I currently have a script that will combine multiple csv files into one, the script works fine except that we run out of ram really quickly when larger files start being used.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
